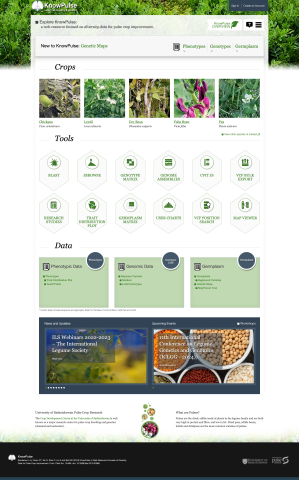
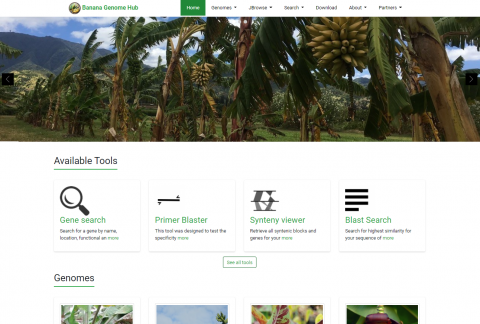
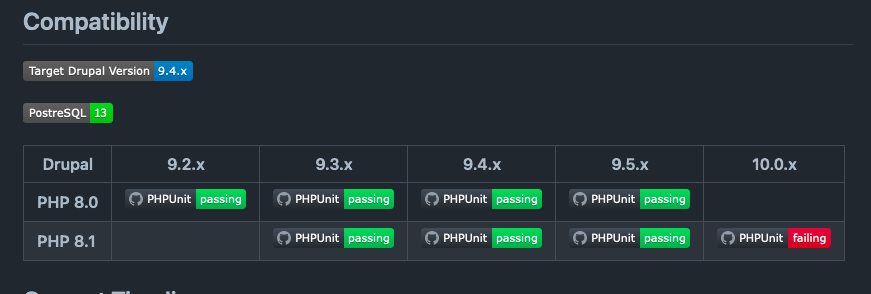
Tripal is a toolkit for construction of online biological (genetics, genomics, breeding, etc), community database web portals, and is a member of the GMOD family of tools. Use Tripal out-of-the box to create a basic genomics site (with no programming) or customize using Tripal's Application Programming Interface (API). Tripal is free and open-source (GNU General Public License version 2), allows for extensive customization and is backed by a helpful user community. A strength of Tripal is our community of developers. Customization and extension of Tripal can be created and shared with other sites as modules allowing you to create your own tools and visualizations or leverage those developed by groups around the world. To see what features Tripal provides, see the Tripal User's Guide and the Developer's Guide! If you have any questions, thoughts, or concerns, we would love to hear from you on our GitHub Issue Queue!
Sites Using Tripal
Tripal sites are found worldwide.
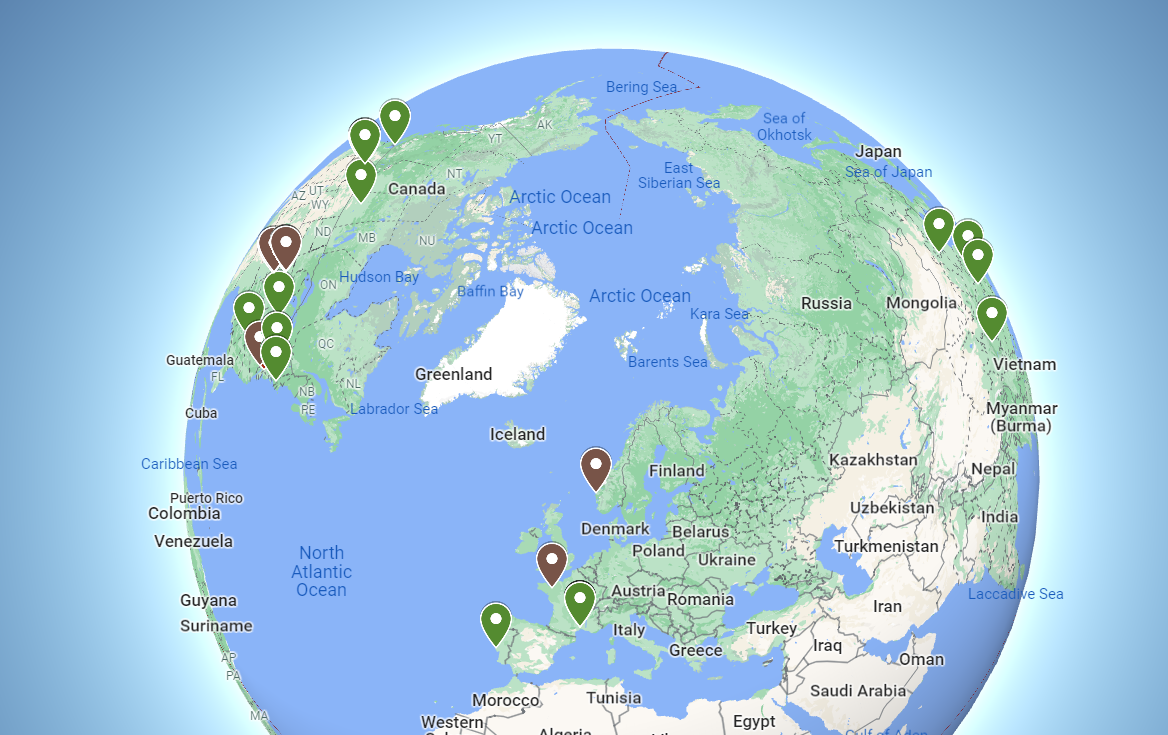
Browse the selected sites below or view more
News
This release represents a major update to Tripal v4. Site developers can now fully publish all content types. This release should allow all Tripal v3 sites managers to begin exploration for how to upgrade their Tripal v3!
For registration and abstract submission please see the Events Page.
The Main Lab Bioinformatics team is happy to announce the release of Tripal MegaSearch v1.4.0 https://gitlab.com/mainlabwsu/tripal_megasearch/
Technical Blog Posts
The Helium Visualization Framework is a visualization tool for large-scale plant pedigree data with the option to overlay categorical data. We developed the Helium Data Exporter module to transform germplasm and raw phenotypic data from a Tripal web portal into a compliant format for seamless and error-free data conversion. Together, this module and Helium provide meaningful assistance to plant breeders and researchers to predict and visualize inheritance.